Question
DNA Strand DNA, or deoxyribonucleic acid, is the primary carrier of genetic information in most organisms. The information in DNA is represented using a string
DNA Strand
DNA, or deoxyribonucleic acid, is the primary carrier of genetic information in most organisms. The information in DNA is represented using a string of nucleotides. There are four kinds of nucleotides: Adenine, Thymine, Cytosine, and Guanine. They are typically abbreviated to their initials, so a strand of DNA might be expressed as ACTTGAT.
These strands join together with a complementary strand to form DNAs famous double helix shape. Adenine and Thymine always pair up and Cytosine and Guanine always pair up. Thus, the strand: ACTTGAT Would always have a complementary strand: TGAACTA Where A in the original lines up with T in the new strand, C in the original matches G in the new strand, etc

----------------------------------------------------------------------------------
DNAStrand.h
#ifndef DNAStrand_H #define DNAStrand_H
#include
class DNAStrand { public: DNAStrand(const char* startingString);
DNAStrand(const DNAStrand& other);
DNAStrand& operator=(const DNAStrand& other);
~DNAStrand();
bool operator==(const DNAStrand& other) const;
char operator[](int index) const;
DNAStrand operator+(const DNAStrand& other) const;
int getLength() const;
DNAStrand getComplement() const; DNAStrand substr(int start, int length) const;
private: DNAStrand(int length);
char* bases = nullptr; ///Tracks array contining a char array int length = 0; ///Length of the char array and DNAStrand };
std::ostream& operator
#endif
-----------------------------------------------------------
DNAStrandTester.cpp
//Bring in unit testing code and tell it to build a main function #define DOCTEST_CONFIG_IMPLEMENT_WITH_MAIN //This pragma supresses a bunch of warnings QTCreator produces (and should not) #pragma clang diagnostic ignored "-Woverloaded-shift-op-parentheses" #include "doctest.h"
//Use Approx from doctest without saying doctest::Approx using doctest::Approx;
#include
//Include your .h files #include "DNAStrand.h"
using namespace std; TEST_CASE( "Basic Constructor, getLength" ) { cout
DNAStrand str1("ACGTAGCT"); REQUIRE( str1.getLength() == 8 );
DNAStrand str2("AACC"); REQUIRE( str2.getLength() == 4 ); }
TEST_CASE( "Basic Constructor, ==" ) { cout
//Verify constructor actually stores the correct data DNAStrand str1("ACGTAGCT"); //If you put the == inside the REQUIRE it will not compile without
DNAStrand str2("AACC"); bool isMatch2 = (str2 == DNAStrand("AACC")); bool isMatch3 = (str2 == DNAStrand("AAGC")); REQUIRE( isMatch2 ); REQUIRE( !isMatch3 ); }
TEST_CASE( "==" ) { cout
DNAStrand str("AACC"); //Make sure unequal length stands don't cause == issues bool isMatch1 = (str == DNAStrand("AAC")); bool isMatch2 = (str == DNAStrand("AACCT")); REQUIRE( !isMatch1 ); REQUIRE( !isMatch2 );
}
TEST_CASE( "Copy constructor" ) { cout
{ /ew scope DNAStrand str2(str1); //Call copy ctor bool isMatch = (str1 == str2); REQUIRE( isMatch ); //str2 goes away when we leave this block... }
bool isMatch2 = (str1 == DNAStrand("ACGTAGCT")); REQUIRE( isMatch2 ); }
TEST_CASE( "Assignment Operator" ) { cout
{ /ew scope DNAStrand str2("AA"); str2 = str1; //Use assignment op
bool isMatch = (str2 == DNAStrand("ACGTAGCT")); REQUIRE( isMatch ); //str2 goes away when we leave this block... }
//Make sure str1 was not changed bool isMatch2 = (str1 == DNAStrand("ACGTAGCT")); REQUIRE( isMatch2 );
//Verify self-assignment does no harm: { str1 = str1; } bool isMatch3 = (str1 == DNAStrand("ACGTAGCT")); REQUIRE( isMatch3 ); }
TEST_CASE( "operator[]" ) { cout
DNAStrand str2("TTTT"); REQUIRE( str2[0] == 'T' ); REQUIRE( str2[2] == 'T' );
//Check that excpetion is thrown if bad index provided int exceptionsCaught = 0;
try { char c = str2[-1]; } catch (out_of_range e) { exceptionsCaught++; }
try { char c = str2[4]; } catch (out_of_range e) { exceptionsCaught++; }
REQUIRE( exceptionsCaught == 2 ); }
TEST_CASE( "operator+" ) { cout
bool isMatch = (str3 == DNAStrand("AATTCCGG")); REQUIRE( isMatch );
//Check originals aren't broken bool isMatch2 = (str1 == DNAStrand("AATT")); bool isMatch3 = (str2 == DNAStrand("CCGG")); REQUIRE( isMatch2 ); REQUIRE( isMatch3 ); }
TEST_CASE( "getComplement" ) { cout
DNAStrand str3 = str2.getComplement(); //should be back where we started bool isMatch2 = (str3 == DNAStrand("ACTGG")); REQUIRE( isMatch2 ); }
TEST_CASE( "Operator
stringstream result; result
REQUIRE( result.str() == "AAC GT" ); }
TEST_CASE( "substr" ) { cout
DNAStrand str2 = str1.substr(0,1); bool isMatch2 = (str2 == DNAStrand("A")); REQUIRE( isMatch2 );
DNAStrand str3 = str1.substr(0,3); bool isMatch3 = (str3 == DNAStrand("AAC")); REQUIRE( isMatch3 );
DNAStrand str4 = str1.substr(2,4); bool isMatch4 = (str4 == DNAStrand("CCGG")); REQUIRE( isMatch4 );
DNAStrand str5 = str1.substr(2,6); bool isMatch5 = (str5 == DNAStrand("CCGGTT")); REQUIRE( isMatch5 );
int exceptionsCaught = 0;
try { //Check bad start str1.substr(-1,4); } catch (out_of_range e) { exceptionsCaught++; }
try { //Check bad length str1.substr(3, -1); } catch (out_of_range e) { exceptionsCaught++; }
try { //Check start + length invalid str1.substr(3, 6); } catch (out_of_range e) { exceptionsCaught++; }
REQUIRE( exceptionsCaught == 3 ); }
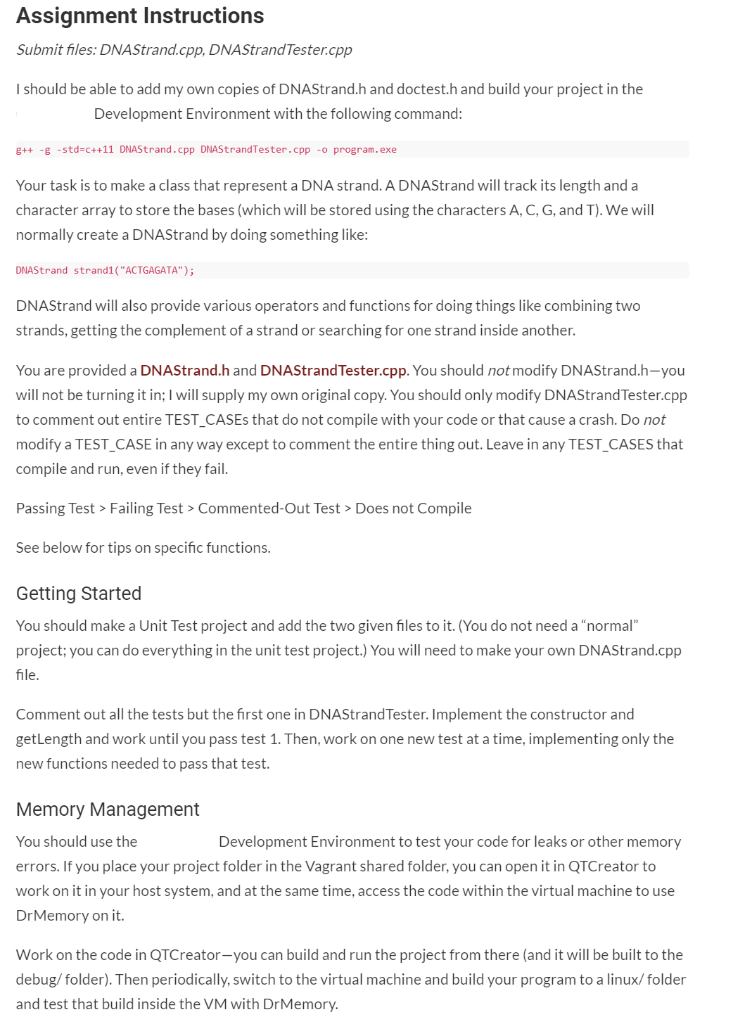
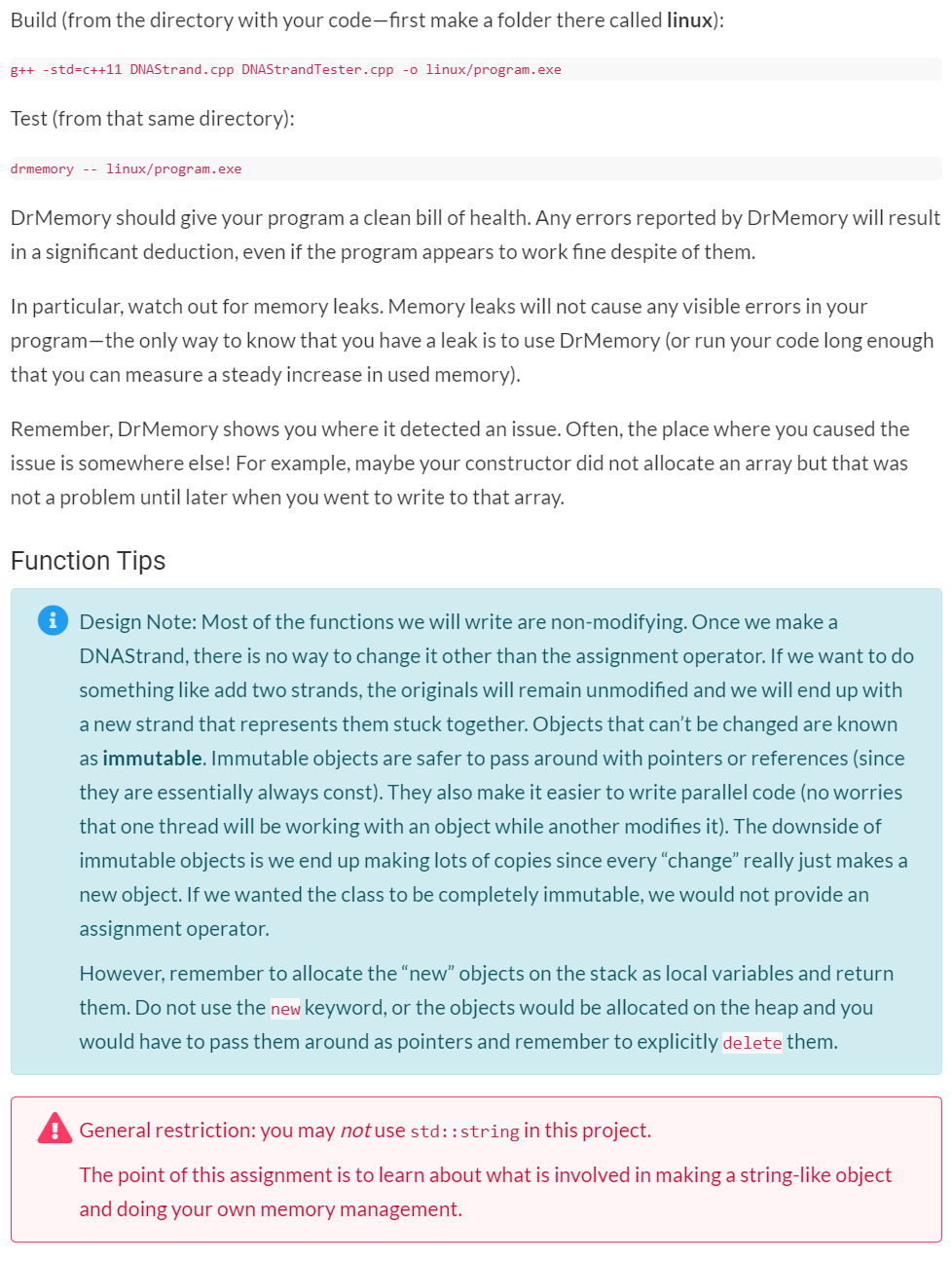
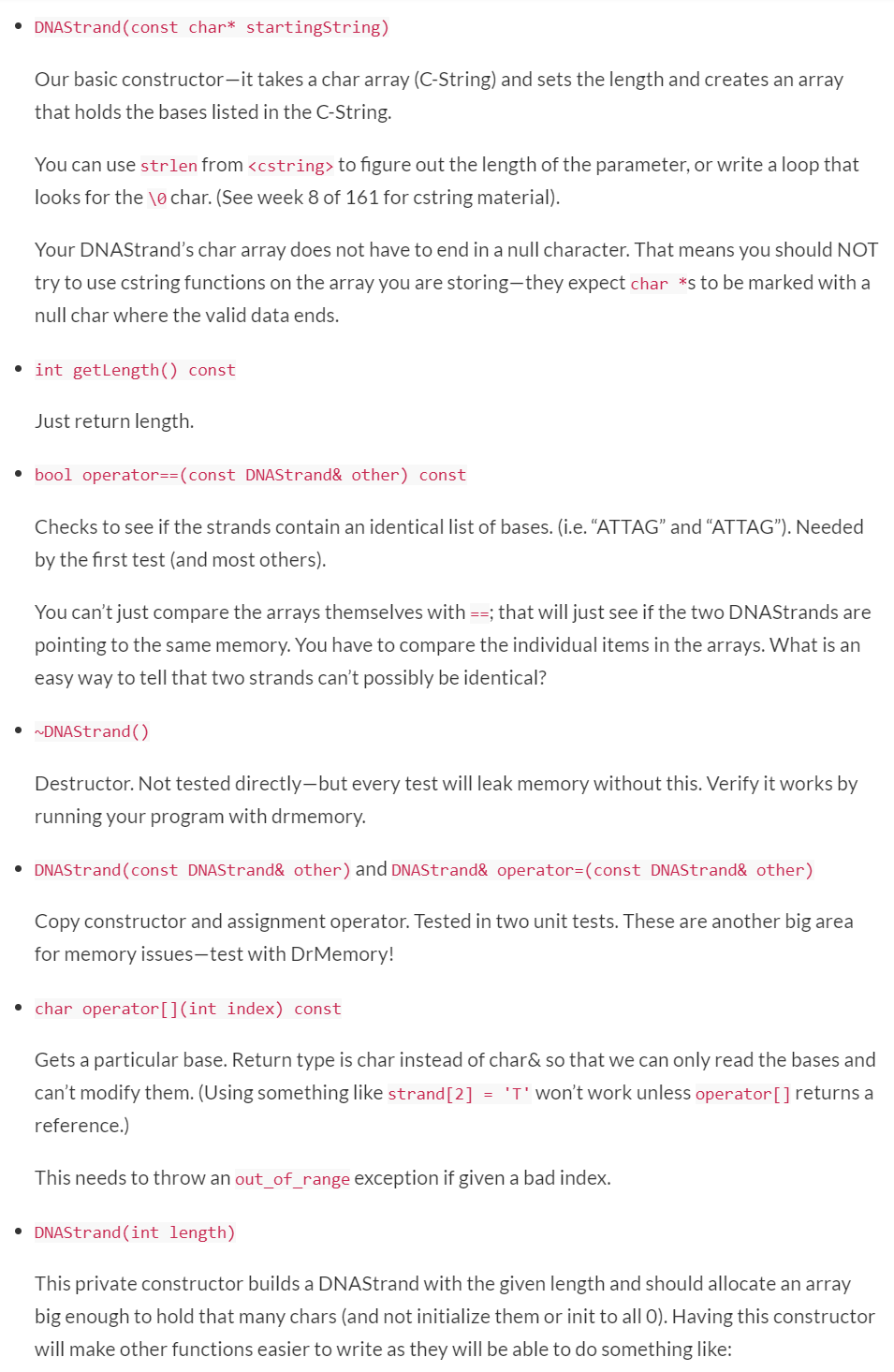
Assignment Instructions Submit files: DNA Strand.cpp, DNAStrandTester.cpp I should be able to add my own copies of DNAStrand.h and doctest.h and build your project in the Development Environment with the following command: g++ -g -std=c++11 DNAStrand.cpp DNAStrandTester.cpp -o program.exe Your task is to make a class that represent a DNA strand. A DNAStrand will track its length and a character array to store the bases (which will be stored using the characters A, C, G, and T). We will normally create a DNAStrand by doing something like: DNAStrand Strandl("ACTGAGATA"); DNAStrand will also provide various operators and functions for doing things like combining two strands, getting the complement of a strand or searching for one strand inside another. You are provided a DNAStrand.h and DNAStrandTester.cpp. You should not modify DNAStrand.h-you will not be turning it in; I will supply my own original copy. You should only modify DNAStrandTester.cpp to comment out entire TEST_CASEs that do not compile with your code or that cause a crash. Do not modify a TEST_CASE in any way except to comment the entire thing out. Leave in any TEST CASES that compile and run, even if they fail. Passing Test > Failing Test > Commented-Out Test > Does not Compile See below for tips on specific functions. Getting Started You should make a Unit Test project and add the two given files to it. (You do not need a "normal" project; you can do everything in the unit test project.) You will need to make your own DNAStrand.cpp file. Comment out all the tests but the first one in DNAStrand Tester. Implement the constructor and getLength and work until you pass test 1. Then, work on one new test at a time, implementing only the new functions needed to pass that test. Memory Management You should use the Development Environment to test your code for leaks or other memory errors. If you place your project folder in the Vagrant shared folder, you can open it in QTCreator to work on it in your host system, and at the same time, access the code within the virtual machine to use DrMemory on it. Work on the code in QTCreator-you can build and run the project from there (and it will be built to the debug/folder). Then periodically, switch to the virtual machine and build your program to a linux/ folder and test that build inside the VM with DrMemory. Build (from the directory with your code-first make a folder there called linux): g++ -std=c++11 DNAStrand.cpp DNAStrandTester.cpp -o linux/program.exe Test (from that same directory): drmemory -- linux/program.exe DrMemory should give your program a clean bill of health. Any errors reported by DrMemory will result in a significant deduction, even if the program appears to work fine despite of them. In particular, watch out for memory leaks. Memory leaks will not cause any visible errors in your program-the only way to know that you have a leak is to use DrMemory (or run your code long enough that you can measure a steady increase in used memory). Remember, DrMemory shows you where it detected an issue. Often, the place where you caused the issue is somewhere else! For example, maybe your constructor did not allocate an array but that was not a problem until later when you went to write to that array. Function Tips i Design Note: Most of the functions we will write are non-modifying. Once we make a DNAStrand, there is no way to change it other than the assignment operator. If we want to do something like add two strands, the originals will remain unmodified and we will end up with a new strand that represents them stuck together. Objects that can't be changed are known as immutable. Immutable objects are safer to pass around with pointers or references (since they are essentially always const). They also make it easier to write parallel code (no worries that one thread will be working with an object while another modifies it). The downside of immutable objects is we end up making lots of copies since everychange really just makes a new object. If we wanted the class to be completely immutable, we would not provide an assignment operator. However, remember to allocate the new objects on the stack as local variables and return them. Do not use the new keyword, or the objects would be allocated on the heap and you would have to pass them around as pointers and remember to explicitly delete them. General restriction: you may not use std::string in this project. The point of this assignment is to learn about what is involved in making a string-like object and doing your own memory management. DNAStrand(const char* startingString) Our basic constructor-it takes a char array (C-String) and sets the length and creates an array that holds the bases listed in the C-String. You can use strlen fromStep by Step Solution
There are 3 Steps involved in it
Step: 1

Get Instant Access to Expert-Tailored Solutions
See step-by-step solutions with expert insights and AI powered tools for academic success
Step: 2

Step: 3

Ace Your Homework with AI
Get the answers you need in no time with our AI-driven, step-by-step assistance
Get Started


