Question
#include #include #include #include ./ppmFile.c #define MAX_VALUE 1000 #define MAX_WIDTH 1680 #define MAX_HEIGHT 1050 struct Point{ float r,g,b; float x,y; }; __constant__ Point p[MAX_VALUE]; __global__
#include
#include "./ppmFile.c"
#define MAX_VALUE 1000 #define MAX_WIDTH 1680 #define MAX_HEIGHT 1050
struct Point{ float r,g,b; float x,y; };
__constant__ Point p[MAX_VALUE];
__global__ void kernel( Point *p, unsigned char *ptr, int sites ) { // Map from blockIdx to pixel position // Your code here ...
// Now find the closest site and color the current pixel accordingly // Hint: In a for loop, measure the Euclidean Distance between the current // pixel and each site. Find the shortest. // Your code here ... }
int getDivisor(int numerator) { int divisor = 31; while((numerator % divisor) != 0) {divisor--;} return divisor; }
int main(int argc, char **argv) {
if(argc
// 1. Obtain width, height of the Voronoi Diagram image and the number of sites from command line ------ // Ensure they are within the pre-set max caps
int w = atoi(argv[1]), h = atoi(argv[2]), points = atoi(argv[3]); w = (w
// 2. Allocate device memory for the Voronoi Diagram image ---------------------------------------------- // Use "Julia Set" for reference // Your code here ... unsigned char *d_output; int image_size = w * h * 3; cudaMalloc((void**)&d_output, sizeof(unsigned char*) * image_size);
// 3. Sites -------------------------------------------------------------------------------------------- // 3.1. Allocate device memory for the sites // The type of each site is a structure named "Point" defined at the top of this program // Your code here ...
// 3.2. Allocate host memory for the the sites. // Then for each site, // - assign a random value say rand()%255+50 to its r, g, and b elemment respectively // - assign a random value say rand()%w to its x, where w is the width of the Voronoi Diagram image // - assign a random value say rand()%h to its y, where h is the height of the Voronoi Diagram image // Your code here ...
// 3.3. Copy all sites from host memory to device memory // Your code here ...
// Launch kernel ---------------------------------------------------------------------------------------- // The pre-defined function "getDivisor" comes convenient. // When it's called with w, getDivisor(w), it generates the largest number (
dim3 dimGrid(w/wGrid, h/hGrid); dim3 dimBlock(wGrid, hGrid);
printf(" dimGrid: (%d, %d)", w/wGrid, h/hGrid); printf(" dimBlock: (%d, %d)", wGrid, hGrid); printf(" Resolution: %dx%d", w, h); printf(" Sites: %d ", points);
// Replace the following line with your code ... // kernel>>(... ...);
// Copy device variables to host ---------------------------------------- // Use "Julia Set" for reference // Your code here ...
// Convert image to the ppm formt and free the host memory.---------------------------------------- // Use "Julia Set" for reference // Your code here ...
// Free device memory // Your code here ... }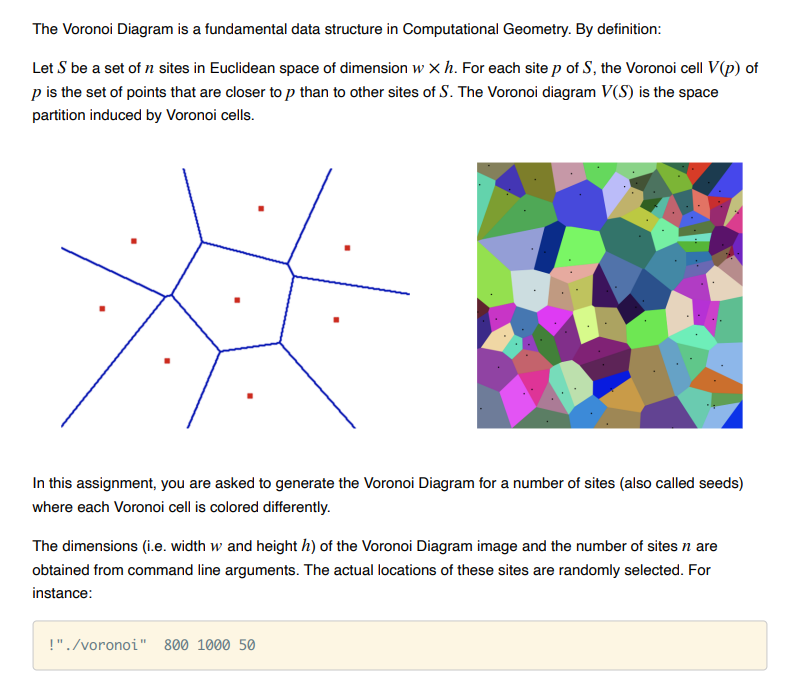
/* * Copyright 1993-2010 NVIDIA Corporation. All rights reserved. * * NVIDIA Corporation and its licensors retain all intellectual property and * proprietary rights in and to this software and related documentation. * Any use, reproduction, disclosure, or distribution of this software * and related documentation without an express license agreement from * NVIDIA Corporation is strictly prohibited. * * Please refer to the applicable NVIDIA end user license agreement (EULA) * associated with this source code for terms and conditions that govern * your use of this NVIDIA software. * */
#include
#include "./ppmFile.c"
#define DIM 1000
struct cuComplex { float r; float i; __device__ cuComplex( float a, float b ) : r(a), i(b) {} __device__ float magnitude2( void ) { return r * r + i * i; } __device__ cuComplex operator*(const cuComplex& a) { return cuComplex(r*a.r - i*a.i, i*a.r + r*a.i); } __device__ cuComplex operator+(const cuComplex& a) { return cuComplex(r+a.r, i+a.i); } };
__device__ int julia( int x, int y ) { const float scale = 1.5; float jx = scale * (float)(DIM/2 - x)/(DIM/2); float jy = scale * (float)(DIM/2 - y)/(DIM/2);
cuComplex c(-0.8, 0.156); cuComplex a(jx, jy);
int i = 0; for (i=0; i 1000) return 0; }
return 1; }
__global__ void kernel( unsigned char *ptr ) { // map from blockIdx to pixel position int x = blockIdx.x; int y = blockIdx.y; int offset = x + y * blockDim.x * gridDim.x; // blockDim.x = 1 so could've been omitted
// now calculate the value at that position int juliaValue = julia( x, y ); ptr[offset*3 + 0] = 255 * juliaValue; ptr[offset*3 + 1] = 0; ptr[offset*3 + 2] = 0; }
int main( void ) { unsigned char *d_output; int image_size = DIM * DIM * 3; cudaMalloc((void**)&d_output, sizeof(unsigned char*) * image_size);
dim3 grid(DIM,DIM); kernel>>(d_output); cudaDeviceSynchronize();
Image *outImage; outImage = ImageCreate(DIM,DIM); ImageClear(outImage,0,0,0);
cudaMemcpy(outImage->data, d_output, image_size, cudaMemcpyDeviceToHost);
const char* outFile = "out.ppm"; ImageWrite(outImage, outFile);
// free(outImage->data);
free(d_output); }
The Voronoi Diagram is a fundamental data structure in Computational Geometry. By definition: Let S be a set of n sites in Euclidean space of dimension wh. For each site p of S, the Voronoi cell V(p) of p is the set of points that are closer to p than to other sites of S. The Voronoi diagram V(S) is the space partition induced by Voronoi cells. In this assignment, you are asked to generate the Voronoi Diagram for a number of sites (also called seeds) where each Voronoi cell is colored differently. The dimensions (i.e. width w and height h ) of the Voronoi Diagram image and the number of sites n are obtained from command line arguments. The actual locations of these sites are randomly selected. For instanceStep by Step Solution
There are 3 Steps involved in it
Step: 1

Get Instant Access to Expert-Tailored Solutions
See step-by-step solutions with expert insights and AI powered tools for academic success
Step: 2

Step: 3

Ace Your Homework with AI
Get the answers you need in no time with our AI-driven, step-by-step assistance
Get Started


