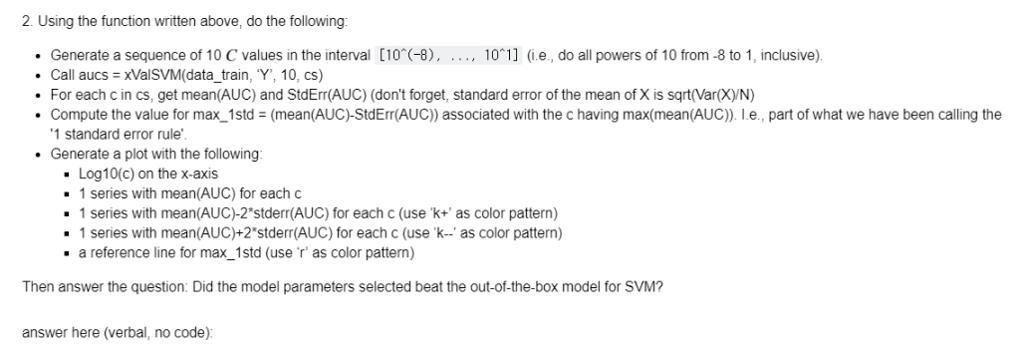

Part 3: Model selection with cross-validation (7 Points) We think we might be able to improve the performance of the SVM if we perform a grid search on the hyper-parameter C. Because we only have 2000 instances, we will have to use cross-validation to find the optimal C Write a cross-validation function that does the following Takes as inputs a dataset, a label name, # of splits/folds ( k ), a sequence of values for C ( cs ) Use sklearn.model_selection.KFold to map each instance to a fold Performs two loops . Outer Loop: for each fold in range(k.) Splits the data into cv-train & cv-val i date according to previously defined fold mappings Inner Loop: for each c in cs o Trains an SVM on training split with C-c, kernel-" inear" o Computes AUC_c_k on validation data o Stores AUC_c_k in a dictionary of values Returns a dictionary, where each key-value pair is: c: [auc.c.1,auc-c_2...auc_c_k] (ie., for each c, we want a list full of auc's from each fold) Note: Use Sklearn's KFold method, but do not use any other cross-validation convenience function. The goal is to learn how to implement the algorithm yourselfl Grading guideline: if this function is done in more than 30 lines (not including empty lines), we will deduct 2 points. In [):\#Don't forget to lapor t any relevant librar les (1.e., sklearn.oross-validation) def xValSVM(dataset, label_name, k, cs): # Code here return aucs Part 3: Model selection with cross-validation (7 Points) We think we might be able to improve the performance of the SVM if we perform a grid search on the hyper-parameter C. Because we only have 2000 instances, we will have to use cross-validation to find the optimal C Write a cross-validation function that does the following Takes as inputs a dataset, a label name, # of splits/folds ( k ), a sequence of values for C ( cs ) Use sklearn.model_selection.KFold to map each instance to a fold Performs two loops . Outer Loop: for each fold in range(k.) Splits the data into cv-train & cv-val i date according to previously defined fold mappings Inner Loop: for each c in cs o Trains an SVM on training split with C-c, kernel-" inear" o Computes AUC_c_k on validation data o Stores AUC_c_k in a dictionary of values Returns a dictionary, where each key-value pair is: c: [auc.c.1,auc-c_2...auc_c_k] (ie., for each c, we want a list full of auc's from each fold) Note: Use Sklearn's KFold method, but do not use any other cross-validation convenience function. The goal is to learn how to implement the algorithm yourselfl Grading guideline: if this function is done in more than 30 lines (not including empty lines), we will deduct 2 points. In [):\#Don't forget to lapor t any relevant librar les (1.e., sklearn.oross-validation) def xValSVM(dataset, label_name, k, cs): # Code here return aucs








